FLEX: Knowledge-Guided Feature Enhancement
A novel framework for robust and fair pathology foundation models
FLEX is a novel framework designed to solve two critical failures in current pathology AI: poor cross-site generalization and demographic bias. By leveraging expert-guided visual and textual knowledge, FLEX disentangles true biological signals from "site-specific signatures," ensuring robust and fair diagnostics across diverse clinical environments.
The Challenge: Shortcut Learning in Pathology
Pathology foundation models often achieve high performance by learning spurious correlations rather than pathology.
-
Site-Specific Signatures: Variations in tissue preparation, staining, and scanning equipment create unique "signatures" for each hospital. Models learn these signatures as shortcuts. They perform well on data from familiar hospitals (In-Domain) but fail catastrophically on data from new sites (Out-of-Domain).
-
Demographic Bias: Models frequently exhibit performance disparities across racial and ancestral groups due to dataset imbalances and biological variations.
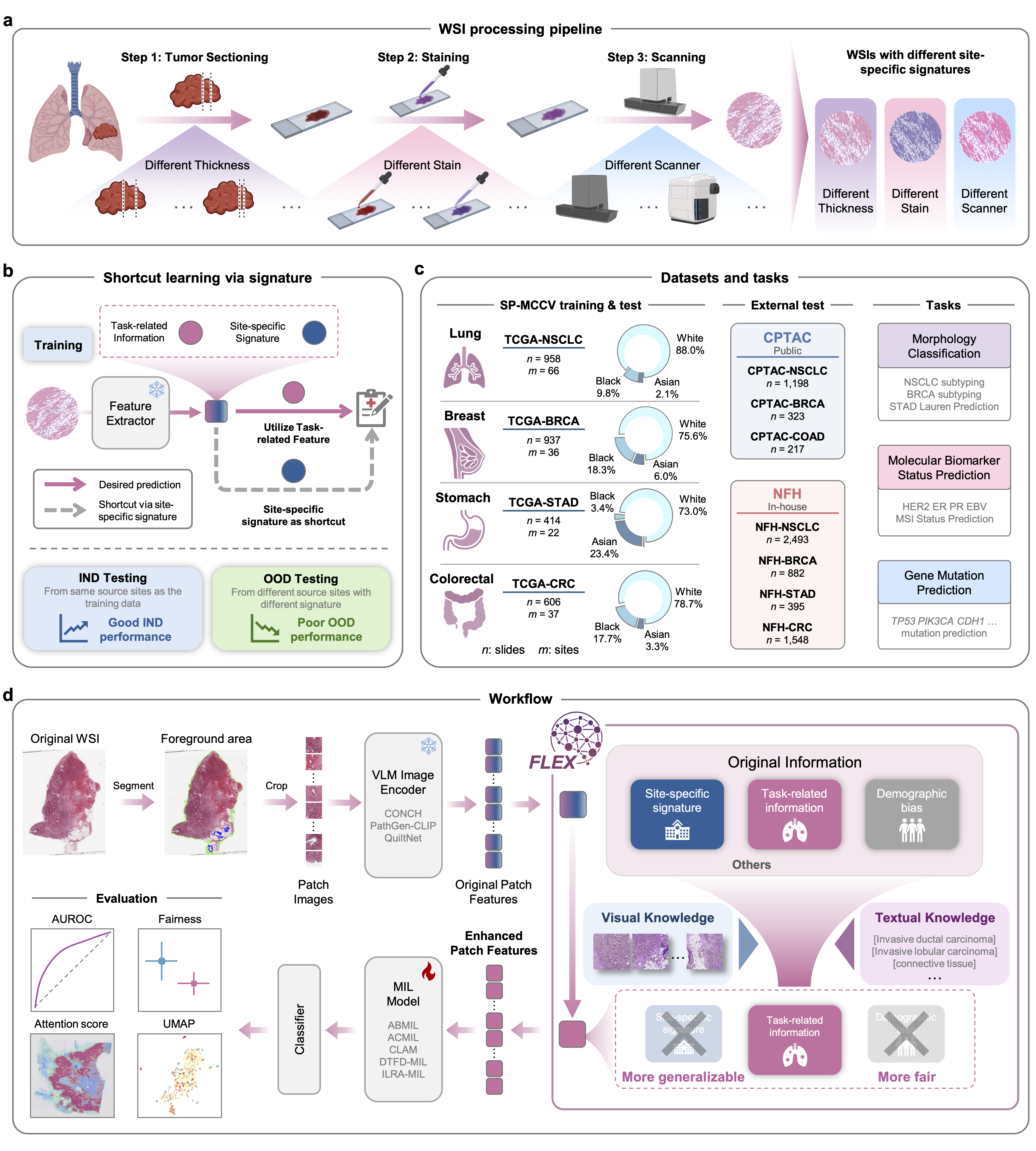 Figure: (a-b) Illustration of site-specific signatures causing shortcut learning. (d) The FLEX workflow utilizing visual and textual knowledge.
Figure: (a-b) Illustration of site-specific signatures causing shortcut learning. (d) The FLEX workflow utilizing visual and textual knowledge.
The Solution: Knowledge-Guided Information Bottleneck
FLEX operates as a model-agnostic module that refines features extracted by Vision-Language Models (VLMs). It employs a Variational Information Bottleneck to filter out noise (site/demographic artifacts) while retaining task-relevant biological information.
The Core Mechanism
FLEX guides feature learning using two forms of expert-verified prior knowledge:
Visual Concepts:
- Representative patch images for specific cancer subtypes are identified by attention mechanisms.
- These patches are filtered by GPT-4o and verified by human pathologists to ensure they represent true biological features.
Textual Concepts:
- Clinically accurate text prompts (e.g., "invasive ductal carcinoma") are generated and approved by board-certified pathologists.
- Why text? Textual descriptions are inherently free of image-based artifacts (like scanner noise or stain variations), providing a clean signal for alignment.
Training Process: An InfoNCE loss aligns patch features with these clean textual concepts, effectively "forcing" the model to unlearn site-specific shortcuts.
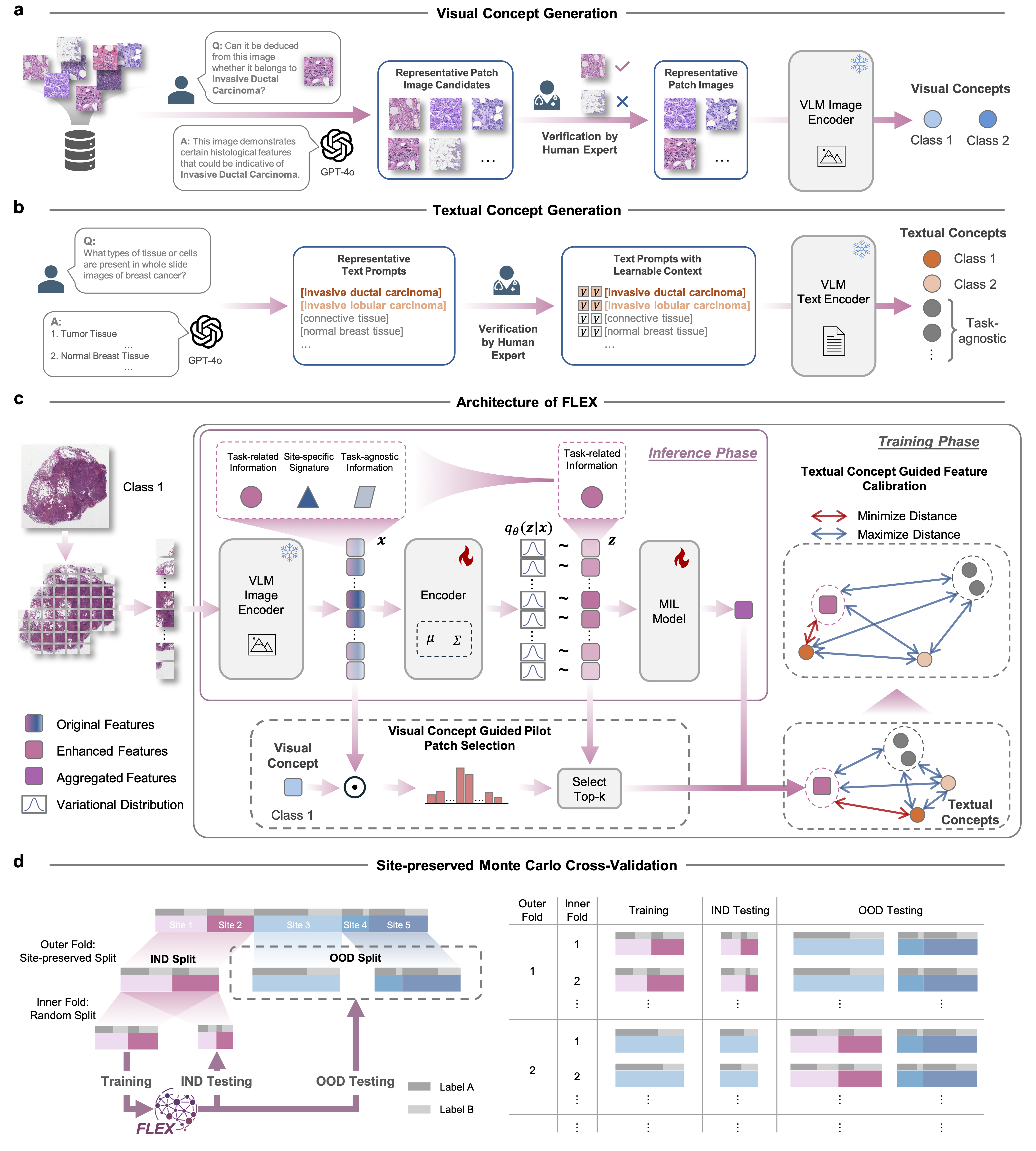 Figure: The architecture of FLEX. (a-b) Generation of visual and textual concepts. (c) The variational information bottleneck guided by concept constraints.
Figure: The architecture of FLEX. (a-b) Generation of visual and textual concepts. (c) The variational information bottleneck guided by concept constraints.
3. Key Results & Evidence
Evaluation was conducted across 16 clinical tasks (morphology, biomarkers, gene mutations) using the TCGA dataset for training, and CPTAC and NFH (in-house) datasets for external validation.
A. Superior Cross-Domain Generalization
FLEX significantly closes the gap between in-domain and out-of-domain performance.
- Robustness: In site-preserved cross-validation, FLEX reduced the performance drop seen in baseline models. For the STAD Lauren task, FLEX increased accuracy on unseen sites by 15.13%.
- Zero-Shot Success: When applied to completely unseen external cohorts (CPTAC and NFH) without fine-tuning, FLEX consistently outperformed baselines (see Figure 2).
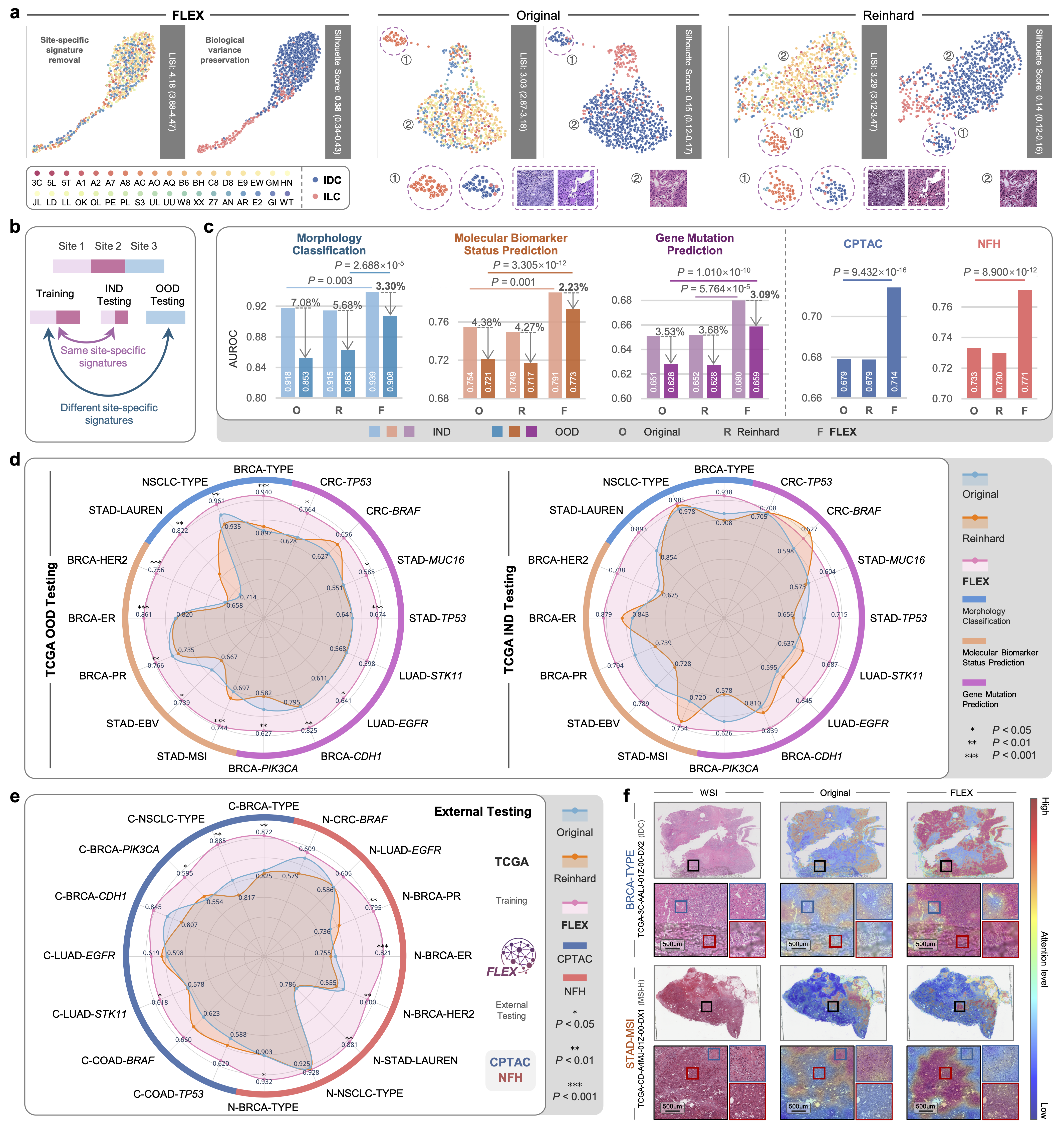 Figure: (a) UMAP shows FLEX removing site clustering. (c) FLEX outperforms baselines in both In-Domain (IND) and Out-of-Domain (OOD) settings.
Figure: (a) UMAP shows FLEX removing site clustering. (c) FLEX outperforms baselines in both In-Domain (IND) and Out-of-Domain (OOD) settings.
B. Improving Demographic Fairness
FLEX creates more equitable diagnostic tools.
- Reduced Disparities: The framework reduced the performance gap between racial groups (e.g., White vs. Black patients) across multiple tasks.
- Equitable Detection: Analysis of True Positive Rate (TPR) disparities shows that FLEX yields diagnostic decisions that are less biased toward specific demographic groups compared to standard stain normalization methods.
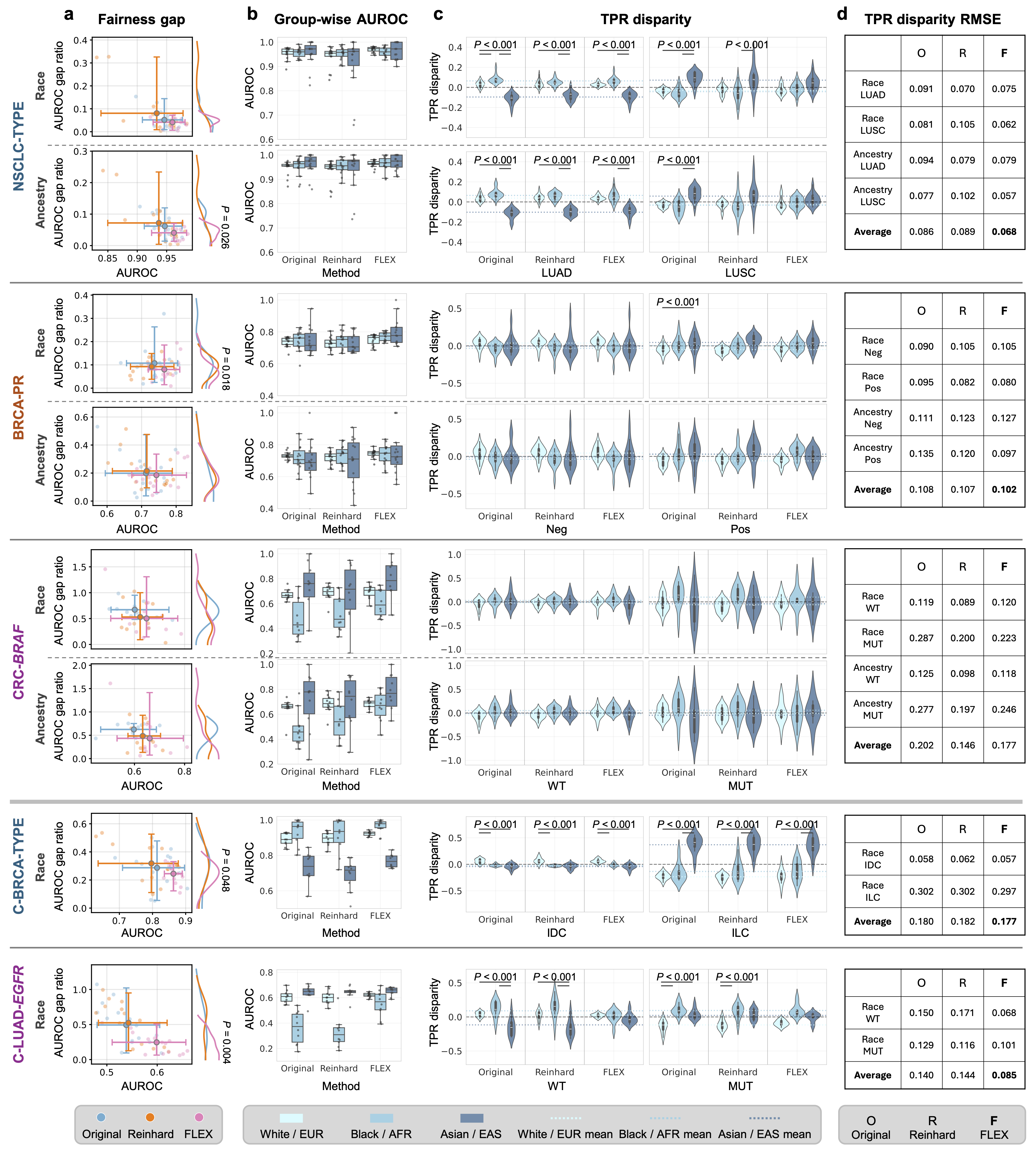 Figure: Evaluation of demographic fairness. FLEX (Pink) consistently shows lower fairness gaps and lower TPR disparity compared to baselines.
Figure: Evaluation of demographic fairness. FLEX (Pink) consistently shows lower fairness gaps and lower TPR disparity compared to baselines.
C. Versatility and Interpretability
- Model Agnostic: FLEX proved effective across different foundation models (CONCH, PathGen-CLIP, QuiltNet) and various MIL architectures.
- Interpretability: Attention maps reveal that while baseline models focus on noise or background tissue, FLEX focuses precisely on tumor regions and relevant histological structures.
4. Conclusion
FLEX establishes a new standard for deploying pathology AI in real-world clinical settings. By strategically using multimodal prior knowledge to suppress site-specific and demographic noise, it offers a pathway to reliable, responsible, and equitable computational pathology.
Citation
@article{huang2025flex,
title={Knowledge-Guided Adaptation of Pathology Foundation Models Effectively Improves Cross-domain Generalization and Demographic Fairness},
author={Huang, Yanyan and others},
journal={Nature Communications},
year={2025}
}
